Welcome to BEable-GPS!
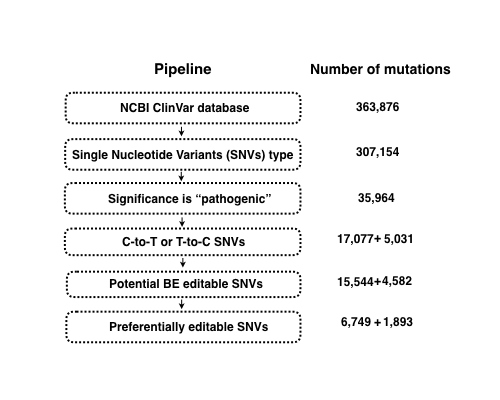
This database is a guidance for the potential editable pathegonic point mutations by CRISPR base editors (BEs). It judges whether the pathogenic mutations collected from NCBI ClinVar database have the potential to be editable by BEs, which is of great importance if these point mutations could be corrected by BEs to cure the diseases or created by BEs to make a disease model.
23/10/19 The paper of BEable-GPS was published in Genome Biology.
30/8/19 The revised version was tested.
22/11/18 BEable-GPS website was tested.
8/10/18 The search function of BEable-GPS website was completed.
- For BEable-GPS:
- Wang Y#,Gao R#,Wu J#,Xiong YC,Wei J,Zhang S,Yang B,Chen J*,Yang L*.Comparison of cytosine base editors and development of the BEable-GPS database for targeting pathogenic SNVs.Genome Biology,218(2019).
- For Base editors(BEs):
- 1.Komor AC,Kim YB,Packer MS,Zuris JA &Liu DR.Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage.Nature,2016:533(7603):420-424
- 2.Nishida K,Arazoe T,Yachie N,Banno S,Kakimoto M,Tabata M, Mochizuki M, Miyabe A,Araki M,Hara KY,Shimatani Z,Kondo A.Science,2016,353(6305):aaf8729
- 3.Koblan LW, Doman JL, Wilson C, Levy JM, Tay T, Newby GA, Maianti JP, Raguram A, Liu DR.Improving cytidine and adenine base editors by expression optimization and ancestral reconstruction. Nat Biotechnol,2018,36(9):843-846
- 4.Kim YB,Komor AC,Levy JM,Packer MS,Zhao KT, Liu DR.Increasing the genome-targeting scope and precision of base editing with engineered Cas9-cytidine deaminase fusions.Nat Biotechnol,2017,35:371-376
- 5.Wang L,Xue W,Yan L,Li X,Wei J,Chen M,W J,Yang B,Yang L,Chen J.Enhanced base editing by co-expression of free uracil DNA glycosylase inhibitor.Cell Res,2017,27:1289-1292
- 6.Hu JH,Miller SM,Geurts MH,Tang W,Chen L,Sun N,Zeina CM,Gao X,Rees HA,Lin Z,Liu DR.Evolved Cas9 variants with broad PAM compatibility and high DNA specificity.Nature,2018,556:57-63
- 7.Nishimasu H,Shi X,Ishiguro Soh,Gao L,Hirano S,Okazaki S,Noda T,Abudayyeh O,Gootenberg J,Mori H et.al.,Engineered CRISPR-Cas9 nuclease with expand targeting space.Science,2018,361:1259-1262
- 8.Li X,Wang Y,Liu Y,Wang X,Wei J,Lu Z,Zhang Y,Wu J,Huang X,Yang L,Chen J.Base editing with a Cpf1-cytidine deaminase fusion.Nat Biotechnol,2018,36:324-327
- 9.Gehrke JM,Cervantes O,Clement MK,Wu Y,Zeng J,Bauer DE,Pinello L,Joung JK.An APOBEC3A-Cas9 base editor with minimized bystander and off-target activities.Nat Biotechnol,2018,36(10):977-982
- 10.Wang X,Li J,Wang Y,Yang B,Wei J,Wang R,Huang X,Chen J,Yang L.Efficient base editing in methylated regions with a human APOBEC3A-Cas9 fusion.Nat Biotechnol,2018,36:946-949
- 11.Grunewald J,Zhou R,Garcia SP,Iyer S,Lareau CA,Aryee MJ,Joung JK.Transcriptome-wide off-target RNA editing induced by CRISPR-guided DNA base editors.Nature,2019,569:433-437
NCBI ClinVar Database: public archive of relationships among sequence variantion and human phenotype