Introduction
Summary
For this module, we will provide a pipeline to analyse spatial transcriptome RNA sequencing dataset from 10X Genomics Visium.
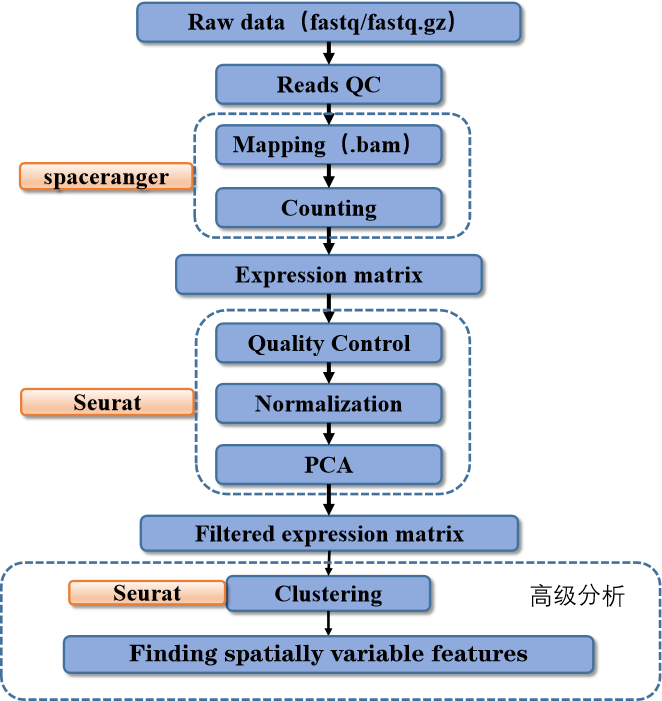
This pipeline starts by reading in the raw fastq data through spaceranger pipeline and return a unique molecular identified count matrix(UMI).
Then we use the count matrix for post preprocessing by Seurat. During post preprocessing, the input count matrix will be filtered, normalized, reduced dimension and clustered by Seurat first.
